Exploratory Data Analysis: Combining Histograms and Density Plots to Examine the Distribution of the Ozone Pollution Data from New York in R
July 29, 2013 9 Comments
Introduction
This is a follow-up post to my recent introduction of histograms. Previously, I presented the conceptual foundations of histograms and used a histogram to approximate the distribution of the “Ozone” data from the built-in data set “airquality” in R. Today, I will examine this distribution in more detail by overlaying the histogram with parametric and non-parametric kernel density plots. I will finally answer the question that I have asked (and hinted to answer) several times: Are the “Ozone” data normally distributed, or is another distribution more suitable?
Read the rest of this post to learn how to combine histograms with density curves like this above plot!
This is another post in my continuing series on exploratory data analysis (EDA). Previous posts in this series on EDA include
- Descriptive statistics
- Box plots
- The conceptual foundations of kernel density estimation
- How to construct kernel density plots and rug plots in R
- Violin plots
- The conceptual foundations of empirical cumulative distribution functions (CDFs)
- 2 ways of plotting empirical CDFs in R
- The conceptual foundations of histograms
Getting the Data and Summary Statistics
Before plotting the histograms and density curves, let’s get the data and calculate the summary statistics. In this post, I add the maximum of the “Ozone” data using the max() function for reasons further explained later.
##### Exploratory Data Analysis of Ozone Pollution Data in New York ##### By Eric Cai - The Chemical Statistician # clear all variables rm(list = ls(all.names = TRUE)) # view first 6 entries of the "Ozone" data frame head(airquality) # extract "Ozone" data vector ozone = airquality$Ozone # sample size of "ozone" length(ozone) # summary of "ozone" summary(ozone) # 3 ways to find number of non-missing values in "ozone" length(ozone[is.na(ozone) == F]) length(ozone[!is.na(ozone)]) n = sum(!is.na(ozone)) # calculate mean of "ozone" mean(ozone) # calculate mean, variance and standard deviation of "ozone" by excluding missing values mean.ozone = mean(ozone, na.rm = T) var.ozone = var(ozone, na.rm = T) sd.ozone = sd(ozone, na.rm = T) var(ozone, na.rm = T) sd(ozone, na.rm = T) max.ozone = max(ozone, na.rm = T)
Kernel Density Plot: A First Non-Parametric Approximation
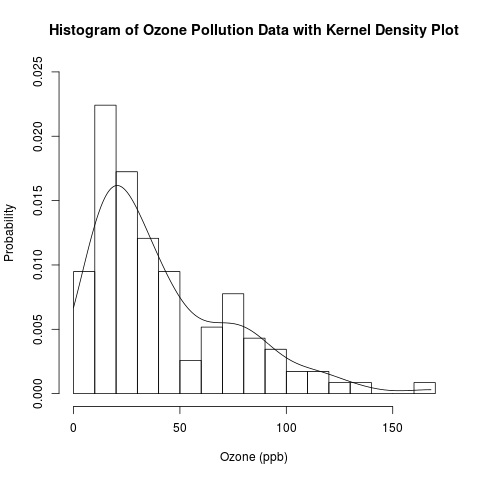
In a previous post, I introduced the conceptual foundations of kernel density estimation. I then wrote a follow-up post to describe the distributions of the “Ozone” data from the “airquality” data set for New York and my own randomly generated ozone pollution data set for a fictitious city called Ozonopolis. It is always a good idea to explore a data set with multiple techniques, especially when they can be done together for comparison. Here is the R code for the histogram with the kernel density plot below it.
- Notice my use of the lines() function to add the kernel density plot.
- Ozone concentration is a non-negative quantity. Thus, I wrote the “from = 0” and “to = max.ozone” options in the density() function to ensure that the support set of the kernel density plot is non-negative.
- Recall that, by default, the density() function uses the Gaussian kernel. You may want to experiment with other kernels, though I did mention in my previous post that the bandwidth is much more important in determining the kernel density estimate.
# histogram with kernel density estimate
png('INSERT YOUR DIRECTORY PATH HERE/histogram and kernel density plot.png')
hist(ozone, breaks = 15, freq = F, xlab = 'Ozone (ppb)', ylim = c(0, 0.025), ylab = 'Probability', main = 'Histogram of Ozone Pollution Data with Kernel Density Plot')
lines(density(ozone, na.rm = T, from = 0, to = max.ozone))
dev.off()
The kernel density plot is a good reference for finding a parametric distribution. If a parametric distribution that fits the data well can be found, inference would be easier. Let’s try to find such a well fitted distribution.
Start with the Normal
The normal distribution is a commonly used distribution for continuous variables with many convenient properties, so let’s try to fit the normal distribution to this data and examine if it is consistent with the histogram.
- Note my use of the dnorm() function to get the normal density values. I used the sample mean and sample standard deviatio as calculated above to define this particular normal distribution.
- I calculated a vector called “ozone.ylim.normal” to get the lowest and largest values of both the histogram and the normal density. I wanted to use it in the y-axis limits of my histogram by setting ylim = ozone.ylim.normal so that the y-values of both the histogram and the normal density curve would be within the vertical limits of the plot. Strangely, the y-axis did not extend to the maximum of the histogram. (Try it yoursefl!) After examining the resulting histogram, I saw that an upper vertical limit of 0.025 would be suitable, so that’s what I used in the ylim option. ozone.ylim.normal was not eventually used for any purpose, but it has been computed here for your own exploration should you wish to do so.
- Note my use of the curve() function to plot this normal density curve.
# histogram with normal density curve
png('INSERT YOUR DIRECTORY PATH HERE/histogram and normal density plot.png')
ozone.histogram = hist(ozone, breaks = 50, freq = F)
ozone.ylim.normal = range(0, ozone.histogram$density, dnorm(ozone, mean = mean.ozone, sd = sd.ozone), na.rm = T)
hist(ozone, breaks = 15, freq = F, ylim = c(0, 0.025), xlab = 'Ozone (ppb)', ylab = 'Probability', main = 'Histogram of Ozone Pollution Data with Normal Density Curve')
curve(dnorm(x, mean = mean.ozone, sd = sd.ozone), add = T)
dev.off()
This normal density curve’s maximum is even lower than the kernel density estimate’s maximum! This is not a very good fit; let’s find another parametric distribution.
The Gamma Distribution?
Notice how the histogram rises quickly for low values of ozone concentration, then decreases gradually. The gamma distribution has this behaviour; let’s give that a try! However, in order to define the gamma function, we need a way to estimate the parameters.
Allow me to parametrize the gamma distribution with the following probability density function (PDF). (There are several ways of parametrizing the gamma distribution, so it is always a good idea to specfiy the PDF for clarity.)
where
.
is the called the shape parameter, and
is called the scale parameter.
Recall that a gamma random variable has the expected value
and the variance
.
Since the sample mean, , and the sample variance,
, are consistent estimators of expected value and variance, respectively, sensible estimates of
and
would be
.
Here is the code for plotting the histogram with the gamma density curve.
# histogram with normal density curve
png('INSERT YOUR DIRECTORY PATH HERE/histogram and normal density plot.png')
hist(ozone, breaks = 15, freq = F, xlim = c(0, 170), ylim = c(0, 0.025), xlab = 'Ozone (ppb)', ylab = 'Relative Frequency', main = 'Histogram of Ozone Pollution Data with Gamma Density Curve')
curve(dgamma(x, shape = mean.ozone^2/var.ozone, scale = var.ozone/mean.ozone), add = T)
dev.off()
This is clearly a better fit than the normal density curve, and it is closer to the kernel density plot, too! (The kernel density plot detects the local maximum at about 75 ppb and has a small “bump” there as a result; the gamma distribution does not have such an accommodation.)


It might also be feasible to fit the histogrammed data with function ‘fitdistr’ of the MASS package and see, which of the available distributions that can be fit delivers the best performance in repsect to lowest residual-sum-squares or AIC.
Cheers,
Andrej
That is a very good suggestion, Andrej! Yes, my method is a visual inspection, but it does help to have some measure of goodness of fit.
Thanks for sharing!
Pingback: Análise Exploratória de Dados com R – Concentração de Ozônio em NYC | Mineração de Dados
Thank you for the positive feedback!
Reblogged this on nishant@analyst.
I’m reading your post in an aim to find a solution of plotting two instgrams in one plot for two dataset that has different size. One data set contains 23 records and the other 200 records. the aim of this is to compare the two dataset and to see if there is a record in the larger dataset that is two standard deviation from the mean of the 23 dataset. But still couldn’t figure this from your article. Am I reading the right article or I’m in the wrong place.
Can you please help me where to start if this is not the right place
Sorry Zainab – this is not the right post for solving your problem. I don’t have any code to do this on my blog.
Good luck!
Eric
Thank you for the informative post. I do have a question regarding your choice to try to see if normal distribution fits your data. It seems like all the observations for the variable are positive (i.e. greater than zero). What is the rationale behind choosing normal distribution, given the fact that the support for normal distribution is from (-infinity, + infinity). Doesn’t Log-normal distribution sound more suitable?
Yes, you are right. The log-normal distribution would be more suitable. However, the normal distribution is good enough for such data. The key is capturing the middle part of the bell curve, especially the mean and the standard deviation.